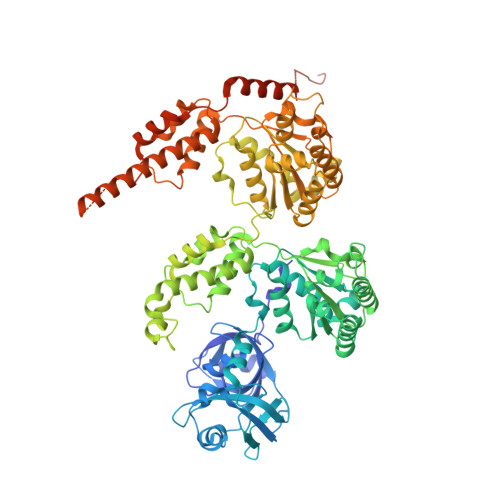
Mechanistic insight into substrate processing and allosteric inhibition of human p97.
Pan, M., Yu, Y., Ai, H., Zheng, Q., Xie, Y., Liu, L., Zhao, M.(2021) Nat Struct Mol Biol 28: 614-625
- PubMed: 34262183 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00617-2
- Primary Citation Related Structures:
7LMY, 7LMZ, 7LN0, 7LN1, 7LN2, 7LN3, 7LN4, 7LN5, 7LN6 - PubMed Abstract:
p97 processes ubiquitinated substrates and plays a central role in cellular protein homeostasis. Here, we report a series of cryo-EM structures of the substrate-engaged human p97 complex with resolutions ranging from 2.9 to 3.8 Å that captured 'power-stroke'-like motions of both the D1 and D2 ATPase rings of p97. A key feature of these structures is the critical conformational changes of the intersubunit signaling (ISS) motifs, which tighten the binding of nucleotides and neighboring subunits and contribute to the spiral staircase conformation of the D1 and D2 rings. In addition, we determined the cryo-EM structure of human p97 in complex with NMS-873, a potent p97 inhibitor, at a resolution of 2.4 Å. The structures showed that NMS-873 binds at a cryptic groove in the D2 domain and interacts with the ISS motif, preventing its conformational change and thus blocking substrate translocation allosterically.
- Department of Biochemistry and Molecular Biology, The University of Chicago, Chicago, IL, USA. panm@uchicago.edu.
Organizational Affiliation: